Newly discovered enzymes that are vital for bacterial defenses can also help chemists characterize the stereochemistry in natural products. The enzymes cut proteins between unusual amino acids, breaking large unwieldy lipopeptides into pieces that are easier to study (J. Am. Chem. Soc. 2026, DOI: 10.1021/jacs.5c17955).
Researchers usually study natural products as molecules secreted by single organisms, but researchers in Pierre Stallforth’s laboratory wondered what happens when microbes cooperate. In 2020, he and his team at the Leibniz Institute for Natural Product Research and Infection Biology identified a pair of bacterial species—a Pseudomonas species and a Paenibacillus species—that work together to survive amoeba attack (Proc. Natl. Acad. Sci. U.S.A. 2021, DOI: 10.1073/pnas.2013759118). When the species are apart, the bacteria fall prey to the amoeba, but when together, they kill their would-be predator.
In a previous study, Stallforth and colleagues had traced the cooperative activity to natural products that the Pseudomonas species produces, including leggy lipopeptides called syringafactins. The Paenibacillus species chops some syringafactins into smaller molecules that are deadly to the amoeba.
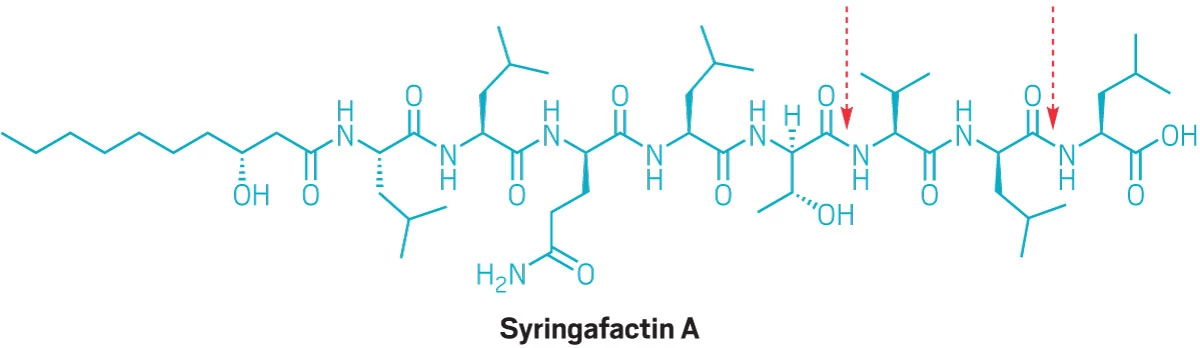
Line structure of the lipopeptide syringafactin A, which features a series of d- and l-amino acid units and several hydrocarbon tails.
Newly discovered enzymes can cut certain bonds (arrows) between specific D- and L-amino acid units in syringafactin A; four other such D-L bonds remain intact.
To analyze the Paenibacillus enzymes at play, Stallforth’s team learned from the mutualistic microbes and did some cooperating of their own: the teamed up with Ute Hellmich and Markus Lakemeyer at the Friedrich Schiller University Jena. In the new study, the researchers exposed Paenibacillus to syringafactins and observed which enzyme-coding genes the microbe expressed more of. This led them to two unusual enzymes behind the bacterium’s defense: DL-peptidases, which break bonds between D- and L-amino acids.
Most amino acids in nature occur in the L-configuration, so LL-peptidases that break the bonds between two L-amino acids are common. In comparison, DL-peptidases are much rarer.
Crucially, the peptidases that the researchers discovered didn’t cleave all the D-L bonds in syringafactins; each recognizes its own D-L bond and leaves the others intact. “We were excited that the peptidases we found were actually quite specific,” says Hellmich. “That makes them valuable.”
Roger Linington of Simon Fraser University, who was not involved in the study, agrees. Existing techniques for analyzing peptides that contain D-amino acids often hydrolyze an entire molecule into its individual amino acids instead, he explains. If a peptide contains the same amino acid in both D and L configurations, this analysis doesn’t show where each occurs. The newly discovered DL-peptidases could “come in handy” by breaking natural product molecules at specific locations, preserving them as larger fragments and revealing that the amino acids on each side of the break are in D and L configurations.
To demonstrate the enzymes’ utility, the three teams modified the peptidases to expand the types of D-L peptide bonds that they could cleave. The researchers also used the enzymes to characterize two natural products—tensin and WLIP, large lipopeptides also produced by Pseudomonas bacteria.
Beyond characterization, the findings may help discover the bioactivity of more natural products, particularly when the molecules have been modified through microbial cooperation. Recognizing that a product natively produced by one organism may not be an active component is important for the natural product community to consider, says Linington.
The research teams are searching for more examples of microbial cooperation that involve natural product modification and that could yield more novel enzymes. “We want to show that this is a ubiquitous phenomenon found in nature,” Stallforth says. Meanwhile, the researchers are also using the DL-peptidases to characterize new natural products—and they hope other research groups will use the peptidases too. “I certainly have it in mind,” Linington says.
Chemical & Engineering News
ISSN 0009-2347
Copyright ©
2026 American Chemical Society