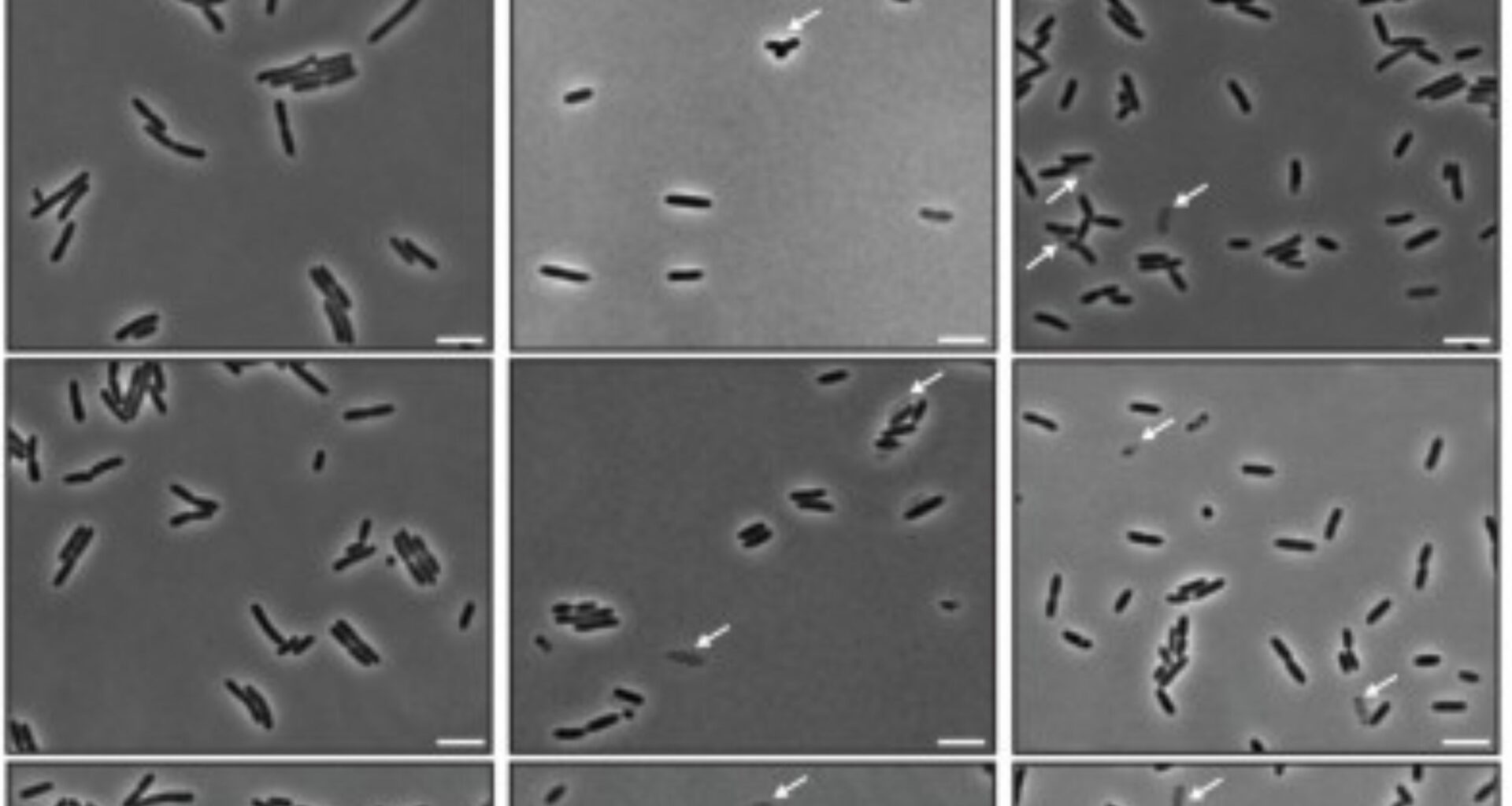
Three unrelated viruses have been shown to kill bacteria by trapping the cell-wall transporter MurJ, which flips essential building blocks for the bacterial wall across the inner membrane, in the same frozen position.
That shared tactic exposes a vulnerable point in one of bacteria’s most essential systems and reframes where new antibiotics could strike.
Within the bacterial inner membrane, MurJ normally flips critical wall-building material to the cell’s outer side, a motion that keeps the protective barrier intact.
By resolving three virus-bound forms of MurJ, Professor William M. Clemons Jr. at the California Institute of Technology (Caltech) demonstrated that distinct viral proteins all clamp onto the same groove and lock the transporter facing outward.
In every case, the protein remains stuck in that outward-facing state, unable to complete the movement required to deliver new wall components.
Such repeated interference at a single site suggests that this exposed position is not incidental, but central to how MurJ can be stopped.
The wall bottleneck
Bacteria stay intact because peptidoglycan, a stiff mesh that gives bacteria shape, keeps cells from bursting, and humans do not make it.
MurJ handles a key wall precursor, moving it from the inner side of the membrane to the outside.
When that transporter stalls, the supply of wall pieces stops at the membrane, and the outer wall never thickens.
“Peptidoglycan is a unique feature of bacteria, and that makes it an attractive antibiotic target,” said Clemons.
Tiny genes, big damage
To burst out of a host, bacteria-infecting viruses must stop wall building fast enough to rupture cells.
Some carry single-gene lysis proteins – small killers encoded by one gene – and researchers call them Sgls.
Instead of making many tools, an Sgl wedges into the membrane and shuts down one essential machine, like MurJ.
That one-hit strategy lets viruses succeed with minimal genetic baggage, and it also points scientists toward drug targets.
Three paths converge
Even though their genomes differ, the three Sgls disable MurJ using the same strategy – an example of convergent evolution.
Despite having no shared sequence, the third lysis protein came from an environmental dataset and still matched the same MurJ groove.
“This is a third genome that evolved a distinct peptide to inhibit the same target in a similar way,” said Clemons.
Repeated hits on the same target suggest this transporter is a weak spot, and other viruses may reveal additional vulnerabilities.
Freezing MurJ in place
To catch MurJ mid-motion, the team used cryo-EM – frozen imaging that maps proteins in fine detail – and saw the gate stuck open.
A pocket inside the transporter normally opens inward to grab a wall precursor, then swings outward to release it.
Each Sgl bound along the groove between two membrane helices, blocking the pivoting movement that powers the flip.
Those cryo-EM maps highlighted small pockets and charged spots on MurJ that could anchor a drug built for fit.
Why exposure matters
For Gram-negative bacteria, microbes with an extra outer membrane, drug molecules often struggle to reach inner wall machinery.
In that stuck-open pose, MurJ opened toward the space between the two membranes, not the cell interior.
Because the pocket faced that gap, a drug could bind without crossing the inner membrane, though the outer barrier remains.
Such access could make MurJ easier to target than hidden enzymes, but any inhibitor still must survive blood and metabolism.
Blueprint for antibiotics
Drug designers can treat the three Sgls as templates, since each one fits MurJ tightly.
Their binding surfaces outline a pocket lined with charged residues, giving chemists clear contact points to target.
Lab screens can now search for small molecules that sit in that pocket, then test whether bacteria stop growing.
Turning that map into a real pill will take years of chemistry, because peptide toxins rarely make safe medicines.
Resistance keeps rising
Nationwide, a Centers for Disease Control and Prevention report estimated that more than 35,000 people die yearly from resistant infections.
“In the U.S. alone, tens of thousands of people die every year from antibiotic-resistant bacterial infections, and that number is rising rapidly,” said Clemons.
Globally, an analysis estimated 1.27 million deaths directly tied to bacterial drug resistance in 2019.
With resistance spreading and new antibiotics arriving slowly, targets highlighted by viruses could keep future infections treatable.
Hidden antibiotic clues
Millions of virus genomes now sit in sequence databases, and many likely encode undiscovered Sgls with bacterial weak spots.
By expressing these tiny genes in bacteria, labs can watch which cells burst, then trace the protein target.
From environmental samples, new viruses can surface without ever being grown, and cryo-EM can still reveal their tactics.
Each time a virus points to a vulnerable bacterial part, drug developers gain another option when older antibiotics fail.
From discovery to treatment
A shared viral solution has now pinned MurJ as a controllable choke point, and its exposed pocket offers a practical blueprint.
Next steps include designing small molecules that hold MurJ in that frozen pose, then testing whether bacteria evolve escape routes.
The study is published in Nature.
—–
Like what you read? Subscribe to our newsletter for engaging articles, exclusive content, and the latest updates.
Check us out on EarthSnap, a free app brought to you by Eric Ralls and Earth.com.
—–