Cells continually decide when to grow, divide, or slow down their activity for proper functioning. A protein called TOR (Target of Rapamycin) plays a vital role in this decision-making.
Found in complex living organisms, TOR acts as a conductor, integrating signals such as nutrient availability, energy levels, and mechanical stress, and adjusting cellular behavior accordingly.
When resources are plentiful, TOR drives cell growth and protein production. Under stress or scarcity, it slows these processes to keep balance. Problems with TOR activity are linked to diseases like cancer and diabetes.
TOR functions through two complexes: TORC1, which is well understood, and TORC2, which is less well understood but vital for cell survival and adaptation.
Observing cellular activity, one molecule at a time
TORC2 is a protein complex that helps keep the cell’s outer membrane stable. It acts like a central hub in feedback loops that control balance. Scientists still don’t fully understand how TORC2 responds when the cell membrane is physically disturbed.
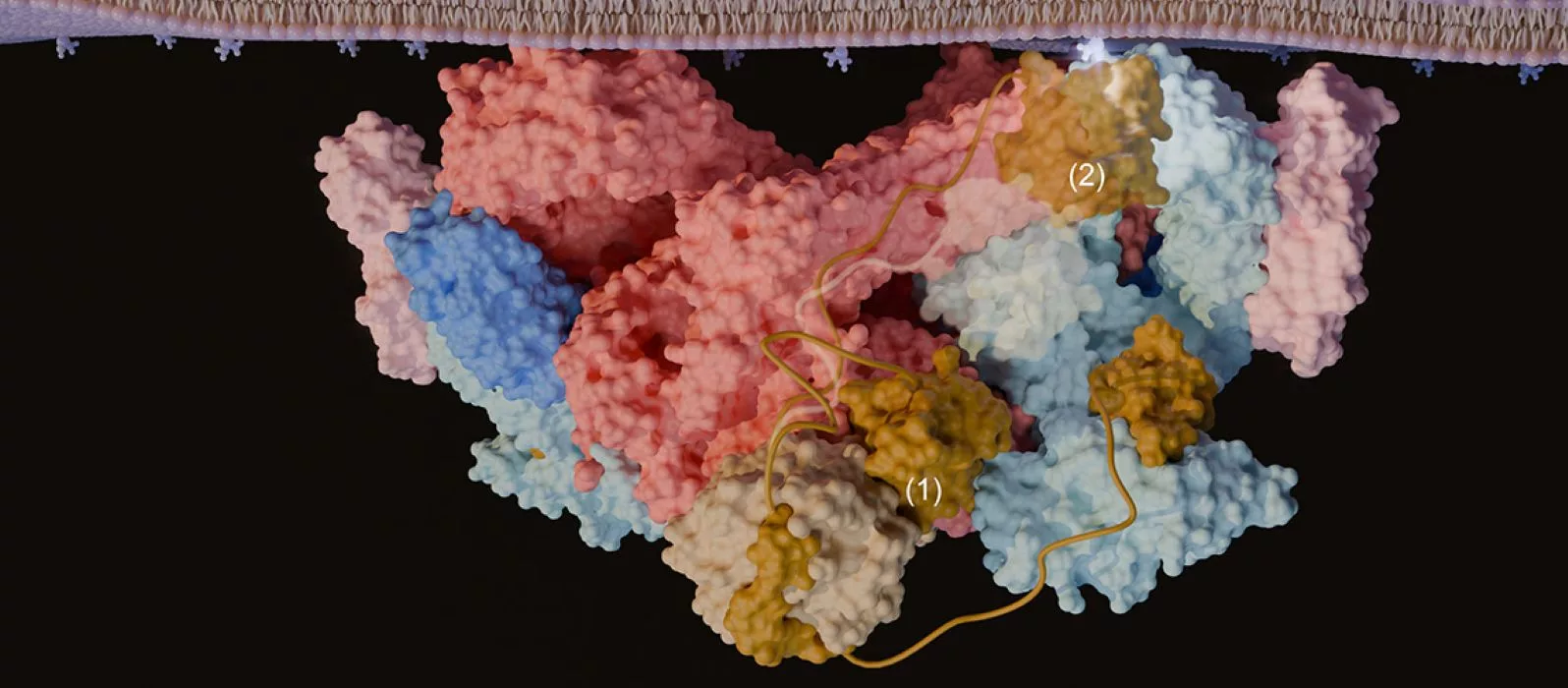
A team from the University of Geneva studied TORC2 in detail using ultra‑high‑resolution imaging. For the first time, they were able to see its structure clearly. They discovered a molecular “cork” that controls how TORC2 is activated at the cell surface. This breakthrough opens the door to new ways to target TORC2, which plays a role in diseases such as cancer and diabetes.
Thanks to the advanced cryogenic electron microscopy facilities at the Dubochet Center in Geneva and Lausanne, researchers were able to observe the biological samples with exceptional precision on the order of an ångström (one ten-billionth of a meter). In fact, the team successfully visualized the structure of TORC2 at up to 2.2 Å resolution.
Because of this approach, researchers could reconstruct the full organization of the complex and revealed the structural elements that had previously escaped observation. The team particularly identified an unexpected regulatory mechanism: part of the protein acts as a true molecular “cork”, blocking the active site of TORC2 and preventing its activation.
Junk DNA could be rewiring our brains
The team also found several protein parts specific to TORC2 that are absent in TORC1. These special domains help TORC2 anchor to the cell membrane and are key to its function.
They found that one subunit (Avo1) inserts into TOR’s active site, giving a way for certain lipids (phosphoinositides) to regulate TORC2. Their experiments also revealed a possible membrane‑binding surface and a charged pocket in another subunit (Avo3) required for TORC2 activation.
Together, the results suggest that signaling lipids activate TORC2 by reshaping its membrane structure, with several conserved subunits working in concert.
Lucas Tafur, a former researcher in Robbie Loewith’s group, said, “These features, previously unseen, represent particularly promising targets for future drugs. Specifically inhibiting TORC2 could make it possible to treat patients who have cancer or diabetes.”
Journal Reference:
Luoming Zou, Maria G. Tettamanti, Caroline Gabus, Ariane Bergmann, Robbie Loewith, and Lucas Tafu. Structural basis for TORC2 activation. Molecular Cell. DOI: 10.1016/j.molcel.2026.03.022