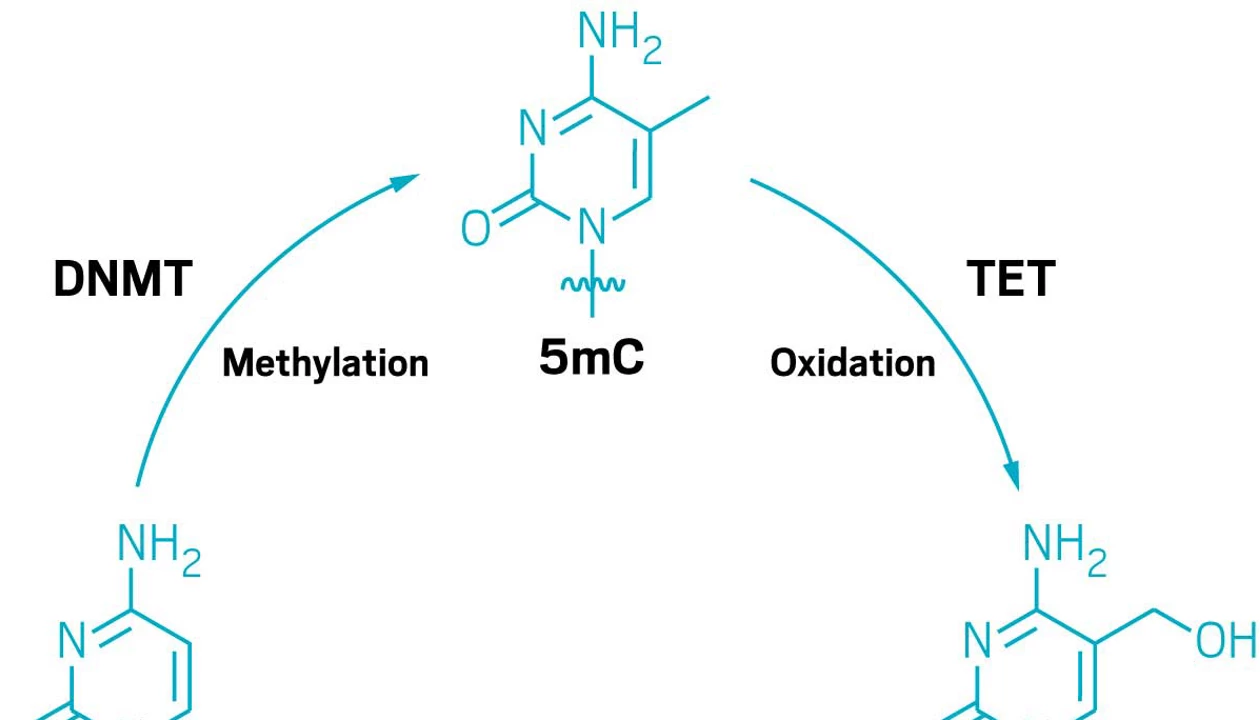
Some researchers say the genetic code has five letters: the four bases and methylated cytosine. The epigenetic marker 5-methylcytosine (5mC) tags parts of the genome to silence their expression, thus defining cells’ identities. But methods to detect 5mC are apt to get it confused with a related modification, 5-hydroxymethylcytosine (5hmC). Researchers have now come up with a sequencing method that can differentiate between the two while also retaining information about the sequences they modify (J. Am. Chem. Soc. 2026, DOI: 10.1021/jacs.5c18450).
In 2009, epigenetics researchers trying to understand how those genes can be turned back on after being silenced by methylation found an enzyme that oxidizes 5mC into 5hmC.
5-Hydroxymethylcytosine showed hints of having biological roles of its own, but it was hard to tell for sure. Bisulfite sequencing, which at the time was used to measure DNA methylation, could not distinguish between the two modifications. “We realized that we had this chemical blind spot,” says Rahul Kohli, a professor at the University of Pennsylvania who led the new study.
In the years since, geneticists have developed several sequencing methods that use enzymes called deaminases, which convert cytosine to uracil but don’t act on 5hmC. Still, deamination comes at a price: turning most C to U effectively reduces the genetic code to three letters. “You lose genetic information to gain the epigenetic information,” chemist Shankar Balasubramanian told C&EN in 2022.
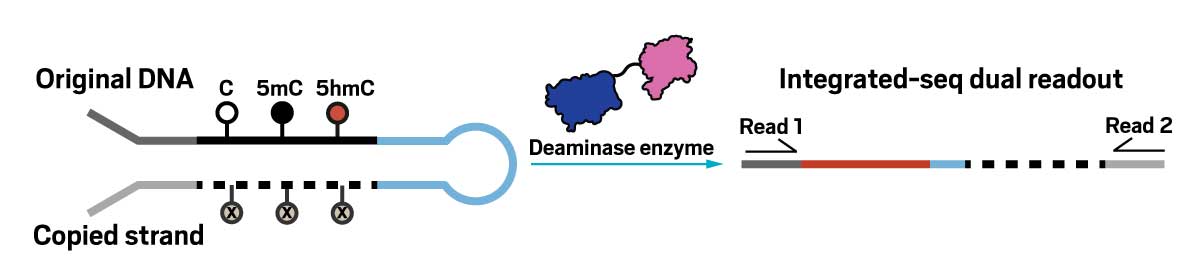
Graduate student Christian Loo worked with Kohli to develop a new method they call integrated sequencing. It involves copying each short DNA sequence into a hairpin duplex, where the cytosine, 5mC, and 5hmC on the new strand are converted into an analog that cannot be deaminated. Then, on the template strand of the hairpin, the researchers deaminate either just unmodified cytosine or both unmodified and 5mC. By sequencing both strands of the duplex, they can recover both the full sequence and its epigenetic markers.
A DNA hairpin molecule with original strand containing C, 5mC and 5hmC, and a copy strand containing an analog at all of those spots. A reaction arrow, with a deaminase enzyme, leads to an unfolded hairpin with two sequencing reads, read1 and read2.
The integrated sequence workflow copies the template DNA and incorporates cytosine analogs that cannot be deaminated. When researchers add a deaminase enzyme, unmodified cytosines are converted to uracil. Sequencing the original (read 1) and copied (read 2) DNA allows researchers to identify all cytosines and their modification status.
Credit:
Courtesy of Rahul Kohli
Being able to extract sequence and modification information from the same molecule is new, Loo says. “There are methods where you can computationally overlay different profiles, but if you have a method that can actually directly link information, that’s incredibly powerful.”
Chunxiao Song, who published a method for differentiating 5mC and 5hmC in DNA extracted from single cells last year (Genome Biol. 2025, DOI: 10.1186/s13059-025-03708-1), tells C&EN in an email that the new method “represents a useful addition to and complement of existing reported and commercial copy-strand–based approaches for DNA modification analysis.”
As for applications, Loo says the team envisions applying the method for cell-free cancer diagnostics. These tests need to find a small number of mutant DNA molecules from cancer cells hidden amid many molecules from healthy cells, and the epigenome of those mutants might give useful information about the tissue of origin.
Laurel Oldach is a senior editor and life sciences reporter at C&EN.
Chemical & Engineering News
ISSN 0009-2347
Copyright ©
2026 American Chemical Society