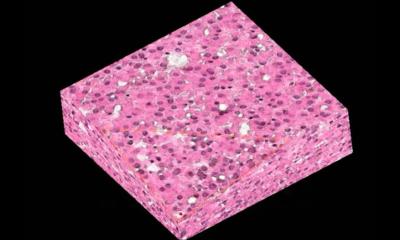
The ability to visualise intact tissue in three dimensions, right down to structures as small and as important as a single cell, opens up so many possibilities in biomedical research and clinical practice
Michela Esposito
The device works by passing X-rays through a small block of tissue while rotating it and capturing thousands of images from different angles, which are then combined using computer reconstruction to create a detailed three-dimensional map of the tissue’s internal structure.
The research team, led by Dr Michela Esposito and Professor Sandro Olivo (both UCL Medical Physics and Biomedical Engineering), say the approach could dramatically simplify histopathology lab work by removing the need for the tissue slicing and staining process, which requires specialised equipment and a range of chemicals.
Professor Olivo said: “Until now, similar types of detailed 3D tissue images were largely only possible at highly specialised research facilities called synchrotrons. These are vast particle-accelerator complexes often the size of football stadiums that cost around £500 million to build and operate, with only around 50 worldwide. The new system achieves comparable results using standard X-ray technology, making it far more accessible for hospitals and research centres that use histopathology for almost all diagnosis.”
Beyond producing 3D images, the technique also allows automated extraction of biological features from tissue samples. As part of their study, the team demonstrates the ability to automatically identify and analyse the shape and size of cell nuclei – parameters known to change in diseases including cancer and common infections – which could help improve diagnostic accuracy.
To help pathologists understand the new images, the research team collaborated with Dr Yang Zhou, Dr Claire Walsh and Professor Peter Lee of UCL Mechanical Engineering to develop an artificial intelligence (AI) method that can convert the X-ray scans into virtual volumes resembling the traditional stained slides that are commonly produced using hematoxylin and eosin (H&E) staining. H&E is one aspect of current histopathology used to stain cells a purple and blue colour to identify the pattern and shape of a cell, which can help to diagnose diseases such as cancer.
The AI-generated virtual volumes enable clinicians to view the data in a familiar format while benefiting from the added information provided by a true 3D representation of tissue architecture. Because the approach is non-destructive, the original sample remains intact and available for further testing if needed – an advantage over existing 3D reconstruction techniques that rely on slicing tissue.
Dr Esposito said: “It’s incredibly exciting to see years of technology development evolve into something with real potential to transform clinical practice. The ability to visualise intact tissue in three dimensions, right down to structures as small and as important as a single cell, opens up so many possibilities in biomedical research and clinical practice. It’s rewarding to think that this work may one day help clinicians make faster, more informed decisions and ultimately improve outcomes for patients.”
Researchers say the method could also help several areas of biomedical research, which rely on identifying 3D tissue structures. Many current imaging approaches use light microscopy tools which struggle to see deep into tissue or require chemical treatments that can alter biological structures. X-ray imaging avoids these limitations and can provide quantitative information about tissue architecture across larger volumes.
The team believes that, with further development, the technology could reduce the time and cost associated with diagnostic pathology while offering clinicians richer information about disease.
Source: University College London