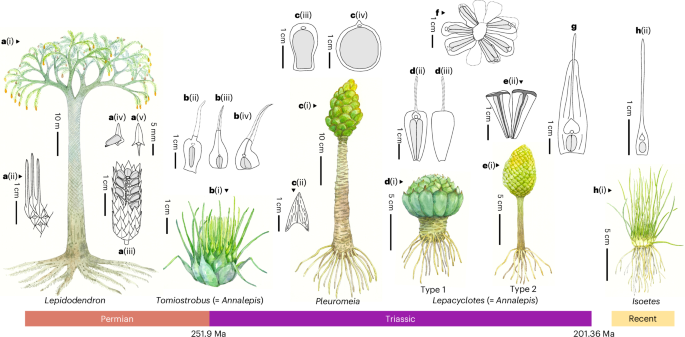
Lycopods sporophyll character identification and measurement
The characters used to differentiate lycopods include root structure, overall plant morphology, cone (strobilus) structure, spore type and number, sporophyll characteristics and sporangium features, incorporating both terminological organ descriptions and topological measurements31. Our study encompasses 127 characters for Isoetales and Lepidodendrales lycopods, with a primary focus on reproductive organs—particularly sporophylls and sporangia—which are more commonly preserved in the fossil record. Although spores are widely used in lycophyte taxonomy, most are found as dispersed specimens rather than in situ, making it difficult to confidently associate them with specific plant macrofossils and resulting in substantial missing data. As the morphology of sporophylls and sporangia is already sufficient to distinguish among taxa in our dataset, we do not emphasize spore data in depth in this study with only simple classification. Future research integrating spore ultrastructures can further refine lycophyte phylogenetic relationships.
Key distinguishing characters include overall plant growth habit (Ch-3), sporophyll phylotaxy (Ch-11), presence or absence of isophylly/heterophylly (Ch-17), apex shape and presence (Ch-58, Ch-59), base shape related to sporophyll attachment (Ch-80), sporangium shape (Ch-88, Ch-89) and sporangium surface ornamentation or structure (Ch-110, Ch-111) (see the loading value of each character in Supplementary Fig. 5). Detailed explanations and figure annotations for these characters are provided in the Supplementary Information. Each specimen of every taxon is coded in a character matrix (Supplementary Data 1), with images and sketches of lycopod sporophyll fossils available in the Supplementary Information.
When selecting characters to distinguish between species, having more characters does not always improve the outcome. Speciation is influenced by isolation and adaptation to different environments. Each species comprises individuals that have evolved under similar environmental conditions, leading to the development of new morphological characters derived from ancestral traits. Therefore, selecting morphological characters with functional role is crucial, especially for studies related to plant physiology. Including too many non-functional characters can dilute the results and reduce their reliability. Characters inherited from common ancestors should be excluded when performing clustering within the same family or order. In addition, random characters lacking functional roles—potentially arising from genetic mutations or preservation biases rather than natural selection—should also be excluded.
In animal phylogenetics, characters are categorized and weighted based on their functional roles69. Similarly, in this study, we have reviewed and discussed the potential functions of the characters used to inform subsequent phylogenetic and ecological analyses. Many characters, such as sporophyll shape and sporangium position, are related to water transport capabilities, while sporophyll base shape affects the attachment and transport of sporophylls on the central axis14,29. Detailed functional inferences for most characters are provided in the discussion section of the Supplementary Information. However, some characters in our matrix lack clearly defined functions, a challenge exacerbated by the limited availability of close extant relatives and the recent extinction of many genera70,71,72. Given the existing gap between plant morphology and function, each character in our matrix is considered equally important71,72.
PCA
Two-dimensional PCA was conducted on the presence/absence of data for lycopod characters, using Euclidean distances in PAST (v4.02)10,71. The method effectively reveals both gradual and distinct variations in sporophyll morphology. Gradual variations are considered within-species diversity, while distinct variations are interpreted as representing different species or subspecies. To capture as much morphological variation (heterophylly) as possible within each taxon, all available plant fossil samples were included in the PCA. In cases where fossils were incomplete but identifiable, missing portions were inferred by comparison with better-preserved specimens of the same species. Fossils that were poorly preserved with unpredictable missing parts or lacking critical information were excluded from the analysis. Consequently, some lycopods of interest may be absent from the dataset. Researchers are encouraged to follow the protocols outlined in the Supplementary Information for incorporating their own fossil collections to enhance the database.
We used original taxonomic names rather than combined or revised names to avoid conflating data and introducing potential biases. For instance, ref. 25 proposed synonymizing dispersed sporophylls previously classified as four species by ref. 30 into a single species, Tomiostrobus sinensis, which was excluded from our analyses.
In the PCA, each character represents an independent dimension, with data point locations determined by their Euclidean distances across these dimensions. Taxa are grouped based on all data points corresponding to a specific species or genus. For visualization, the high-dimensional taxon volumes were projected into a two-dimensional space that captures the maximum amount of character information. The summary scores for each principal component (PC), representing the percentage of variance explained, are listed in Supplementary Table 2. The top three principal components are used for generating the two-dimensional morphospace plots, with the highest score PC1–PC2 shown in Fig. 2 and additional PC1–PC3 plots in Supplementary Fig. 1. Note that only a subset of character information is included in the PCA analysis.
In the PCA plots, polygons of different colours represent clusters of sporophyll characters corresponding to individual species groups. The area of these polygons reflects the range of morphological variation within each taxon, with larger areas indicating greater variation71. Proximity between polygons suggests potential close relationships that warrant further NNA. Overlapping morphospaces are interpreted as potentially representing subspecies. In the PCA analyses shown in Fig. 2, certain characters present or absent in all selected taxa were excluded to prevent data dilution (highlighted as red in Supplementary Data 1). All the fossils are preserved in Room 014B, Main Building, China University of Geosciences (Wuhan).
NNA of cladistic matrix
Neighbourhood network (NNA) is a clustering method that incorporates all characters and is widely used in phylogenetic analysis. It is particularly useful for phylogenetically unsorted taxa, such as most plant fossils, where homoplastic (incompatible) signals can overshadow phylogenetic signals, potentially leading to incorrect tree inferences72. Unlike dichotomous tree methods, neighbourhood network effectively handles non-tree and incompatible signals by representing them as a network, thus providing a more accurate depiction of ancestral–descendant relationships72.
For phylogenetic NNA analyses, we selected one ‘best-preserved’ specimen per taxon to represent the taxon. However, given the heterophylly within taxa as illustrated by the polygon areas in our PCA results, defining the best-preserved fossil can be ambiguous. To minimize subjective bias and mitigate the influence of incompatible data, we selected only one specimen per species that was closest to the centroid of the polygons in the PCA results, reflecting the morphological variation of sporophylls within each lycopod taxon. All taxa and character matrix data were stored in Mesquite (v3.70) and uploaded into PAUP (v4) for distance matrix calculations. The resulting distance matrix was then used to generate the neighbourhood network in SplitTree (v4.18.3). For detailed procedural instructions, refer to ref. 72.
The distance between each tip in the NeighborNet represents the morphological distance between samples, with a 0.1 scale bar indicating the distance in pixels.
We compared the results of the NNA with the PCA results to ensure consistency in phylogenetic information. Both results indicate 12 independent genera of lycophyte sporophyll: Palaeozoic Cyclostigma, Achlamydocarpon, Lepidostrobophyllum, Mazocarpon, Lepidostrobus and Moscovstrobus, and Mesozoic to recent Lycostrobus, Cyclostrobus, Pleuromeia (Pleuromeialean, Lycomeia), Isoetes (Isoetites), Tomiostrobus and Lepacyclotes (Fig. 3). There are clear transitional taxa between each genus in the families Isoetaceae and Pleuromeiaceae. For example, Tomiostrobus (Skilliostrobus) australis (number 310) occurs between Tomiostrobus and Isoetites, while the Lepacyclotes found in North China during the Middle Triassic have the highest similarity with the Permian–Triassic transitional Tomiostrobus angusta. The latter is included within Tomiostrobus zeilleri, and Pleuromeia shaolinii is associated with Pleuromeia and Cyclostrobus (Fig. 3). Based on this comparison, we are able to revise the taxonomy of Triassic Isoetales lycopod sporophylls to robustly distinguish genera, species and subspecies based on our presence–absence data and our morphometric analysis (Supplementary Table 2). Our result suggests there are 26 species including 44 subspecies on a global scale, rather than the 73 species suggested by the existing taxonomy (Supplementary Table 4). Our dataset contains recent Isoetes species and comparable fossil lycopod species, providing a window that links fossil plants to their living descendants. This allows for an exploration of the linkage between morphology, genetics and phylogeny. In Supplementary Table 4, the red and bold taxa are distinct extant species with species designation via either morphological and/or genetic information; thus, these occurrences represent valid taxa and should not be synonymized with taxa in the same branch of the NNA tree. Comparisons between our revised taxonomic groupings based solely on morphology and the current genetically based phylogenetic species lists of Isoetes and Isoetites suggests that our dataset and data processing methods (PCA and NNA) might have artificially reduced the diversity. This is due to factors including (1) morphological characters from other parts of the plant aside from sporophylls distinguishing living species, (2) loss of morphological information during fossilization and (3) the increasing primacy of genomic information in systematics of living species. For example, in our morphological analysis the extant species Isoetes cangae and Isoetes serracarajensis resolve as a single species, whereas molecular analysis identifies them as distinct species73. Overall, these results suggest that our morphology-based phylogeny is, predictably, of lower resolution than a genetic-based taxonomic system, especially in closely related species with similar sporophyll organization. However, this integration of extant and extinct plants into a single phylogenetic framework allows us to pose new questions about the ecophysiology of these extant floras.
Carbon isotopes
Carbon isotope ratios reflect the balance of physiological processes in plants, such as photosynthesis, respiration and transpiration over the lifetime of that tissue. These processes are influenced by atmospheric CO2 pressure, temperature, and local environmental factors such as water availability and salinity. To accurately separate physiological differences in palaeo-plants from environmental influences, sedimentary facies analysis is crucial before selecting plant samples for carbon isotope testing.
Over the past decade, we have conducted sedimentary surveys in South China with the assistance of numerous collaborators. We collected plant fossils from various sedimentary facies and reconstructed plant habitats based on fossil preservation conditions and sedimentary facies5. Our study covers floras collected from terrestrial, paralic and deep-sea facies5. To assess the impact of atmospheric CO2 pressure, we sampled plant fossils with carbon films or cuticles from the Late Permian to the Middle Triassic, alongside proxy-based atmospheric CO2 content reconstructions for each substage. The age, facies and palaeoenvironments of each flora are detailed in ref. 5. The specific parts of the plant fossils that were sampled are detailed in the Supplementary Information.
To ensure that the carbon isotope samples are derived from plant fossils, we analysed both the organic matter from the plant fossils and the surrounding rock matrix. An ~1‰ difference in δ¹³Corg between the plant fossils and the surrounding matrix confirms the reliability of the samples54. Matrix samples were cleaned with compressed air to prevent cross-contamination and are documented in the Supplementary Information. Only identifiable plant fossils were sampled. Samples were extracted using an alloy scalpel, with a minimum of 20 mg per plant body fossil and 5 g for surrounding rock. To avoid contamination by surrounding matrix to the plant fossil samples, we systematically scratched as thin layers as possible. To get enough sampling amount for small plants, for example, the Triassic lycopods, some samples of the same species and specimen were gathered as one sample, resulting in fewer samples but higher accuracy of each datapoint. Considering each part of the plants may bear slightly different carbon isotope value, all parts of each plant fossil were sampled, including leaves (vegetative/sporophylls), branches, seeds, petioles and veins.
To eliminate the influence of inorganic carbon on the carbon isotope signal, all samples for organic carbon isotope testing—including plant carbon such as cuticles and surrounding rock—were treated with 15% HCl acid then repeatedly rinsed with deionized water before drying at 45 °C and subsequent crushing. The description of each sample and the carbon isotope data are presented in the Supplementary Information. The prepared samples were analysed for organic carbon isotope ratios using a Mat253 Plus (Thermo Fisher, MAT 253 Plus Isotope Ratio Mass Spectrometer) and a Delta V advantage (Thermo Fisher) at the State Key Laboratory of Biogeology and Environmental Geology, China University of Geosciences (Wuhan), and an EA-IRMS system (Elemental Analyzer–Isotope Ratio Mass Spectrometry) at the Stable Isotope Facility, Department of Plant Sciences, University of California, Davis. For the Mat253 Plus, calibration was based on GBW (Guobiao Wuzhi, Chinese National Standard Reference Materials) standards (GBW04407, −22.43; GBW04408, −36.91‰) with ACET (acetanilide) (−26.33‰) as the internal standard. For the Delta V Advantage, reference materials included USGS40 (−26.39‰), USGS24 (−16.05‰), and IVA33802174 Urea (−37.32‰). For the EA-IRMS system, multiple laboratory reference materials were used for scale normalization and quality control, including caffeine (δ¹³C −34.90 ± 0.09‰; δ¹⁵N −2.74 ± 0.10‰), glutamic acid (δ¹³C −10.98 ± 0.10‰; δ¹⁵N −8.54 ± 0.08‰), glutathione (δ¹³C −18.27 ± 0.07‰; δ¹⁵N −5.00 ± 0.04‰), scallop (δ¹³C −16.74 ± 0.10‰; δ¹⁵N 9.37 ± 0.06‰) and nylon powder (δ¹³C −24.90 ± 0.05‰; δ¹⁵N −1.12 ± 0.16‰), among others. Analytical precision was better than ±0.1‰ (1σ) for standards and typically within ±0.2‰ for samples, with a maximum uncertainty of ±0.5‰ in cases of low signal intensity or abnormal matrices (for example, high halogen or sulfur contents). Replicate analyses of samples yielded reproducibility better than ±0.2‰ (Supplementary Table 5). All remaining samples and plant fossils are stored in Room 014B, Main Building, China University of Geosciences (Wuhan) and University of Leeds.
HadCM3BL model simulations
HadCM3BL is an Earth system model that incorporates atmosphere, ocean, land and biosphere, developed by the UK Metoffice and University of Bristol3. Specifically, we use HadCM3LB-M2.1aD with a grid resolution of 3.75° × 2.5° in longitude × latitude in both the atmosphere (19 vertical levels) and ocean (20 vertical levels), using the Arakawa B-grid scheme. The model uses a dynamic vegetation scheme, which is crucial for such studies: the Top-Down Representation of Interactive Foliage and Flora Including Dynamics with the MOESE 2.1 land surface scheme. Desert soil albedo is interactively updated on the basis of the soil carbon content, where low soil carbon concentrations result in a modified soil albedo of 0.32 (average modern-day Saharan albedo).
Typically, the ozone distribution is prescribed as a static latitude–pressure–time distribution in many climate models. However, as the climate warms, the tropopause rises, meaning that stratospheric ozone penetrates into the troposphere, which is unphysical if a pre-industrial tropopause height is prescribed for warm time periods. Instead, the ozone distribution is prescribed using a dynamic approach in which ozone is dynamically coupled to the model tropopause height with constant values for the troposphere (0.02 p.p.m.), tropopause (0.2 p.p.m.) and stratosphere (5.5 p.p.m.). This change makes a negligible difference to the global mean surface temperature but does have a small impact on the stratospheric temperature and winds.
A range of boundary conditions are required to configure the model for Permo-Triassic conditions. The Getech Plc. palaeogeography (land–sea distribution, bathymetry, topography) is used as well as time-specific atmospheric pCO2 (detailed below) and solar luminosity. Each simulation was fully equilibrated in both the atmosphere and deep ocean following a three-stage spin-up protocol so that each simulation is fully equilibrated: (1) the globally and volume-integrated annual mean ocean temperature trend is less than 1 °C per 1,000 years, (2) trends in surface air temperature are less than 0.3 °C per 1,000 years, and (3) net energy balance at the top of the atmosphere, averaged over a 100 year period at the end of the simulation, is less than 0.25 W m−2. These simulations have generally been run for over 10,000 model years to ensure complete Earth system equilibrium. Climate means were then produced from the last 100 years of the simulation.
Using systematic proxy data, including sea surface temperature, atmospheric CO2 and sedimentary observations such as climatically sensitive minerals/facies, HadCM3BL successfully established robust simulations across the PTME interval that shows a mega-El Niño and stronger temperature fluctuations both on land and in the ocean due to the collapse of meridional overturning circulation and a contracted Hadley cell3.
In this work, we ran the end-Permian Changhsingian, PTT and Early Triassic Induan scenarios using HadCM3BL with atmospheric CO2 concentrations of 412 p.p.m., 2,568 p.p.m. and 4,000 p.p.m., respectively, derived from boundary values reconstructed using plant stomatal, palaeosol and plant carbon isotope fractionation proxies4,17,18. All simulations can be found on the Bristol BRIDGE website (https://www.bristol.ac.uk/geography/research/bridge/). After stabilization of the atmosphere–ocean–vegetation coupling, we outputted the average and absolute maximum daily land surface temperature.
The global average maximum daily land surface temperature is the average of each day’s highest temperature over a year, describing the overall thermal intensity experienced by the land surface74. This metric is essential in capturing the cumulative effect of heat extremes, which are critical for assessing the habitability of terrestrial environments, especially for vegetation. Unlike mean annual temperature, this index reflects both the frequency and intensity of high-temperature events, providing insights into seasonal thermal stress and potential physiological thresholds for plant survival and function74.
The global absolute maximum daily land surface temperature, by contrast, captures the single highest temperature recorded in each grid cell of each scenario. This metric reflects the most extreme thermal event experienced at each location, providing crucial information on the upper thermal limits of the environment. It is particularly valuable for evaluating the survivability of organisms under short-term extreme heat stress, which may exceed physiological thresholds even if average conditions are tolerable. This parameter helps identify thermal hotspots and assess the risks of episodic temperature extremes that can drive ecological collapse or restrict species distributions.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.