Ethics declarations
Patients and control participants in FinnGen provided informed consent for biobank research, based on the Finnish Biobank Act. Alternatively, separate research cohorts, collected before the Finnish Biobank Act came into effect (in September 2013) and the start of FinnGen (August 2017), were collected based on study-specific consents and later transferred to the Finnish Biobanks after approval by Fimea (Finnish Medicines Agency), the National Supervisory Authority for Welfare and Health. Recruitment protocols followed the biobank protocols approved by Fimea. The Coordinating Ethics Committee of the Hospital District of Helsinki and Uusimaa (HUS) approved the FinnGen study under ethics statement HUS/990/2017. The study is also approved by the Finnish Institute for Health and Welfare (permits THL/2031/6.02.00/2017, THL/1101/5.05.00/2017, THL/341/6.02.00/2018, THL/2222/6.02.00/2018, THL/283/6.02.00/2019, THL/1721/5.05.00/2019 and THL/1524/5.05.00/2020); the Digital and Population Data Services Agency (permits VRK43431/2017-3, VRK/6909/2018-3 and VRK/4415/2019-3); the Social Insurance Institution of Finland (permits KELA 58/522/2017, KELA 131/522/2018, KELA 70/522/2019, KELA 98/522/2019, KELA 134/522/2019, KELA 138/522/2019, KELA 2/522/2020 and KELA 16/522/2020); Findata (permits THL/2364/14.02/2020, THL/4055/14.06.00/2020, THL/3433/14.06.00/2020, THL/4432/14.06/2020, THL/5189/14.06/2020, THL/5894/14.06.00/2020, THL/6619/14.06.00/2020, THL/209/14.06.00/2021, THL/688/14.06.00/2021, THL/1284/14.06.00/2021, THL/1965/14.06.00/2021, THL/5546/14.02.00/2020, THL/2658/14.06.00/2021 and THL/4235/14.06.00/2021); Statistics Finland (permits TK-53-1041-17, TK/143/07.03.00/2020 (earlier TK-53-90-20), TK/1735/07.03.00/2021 and TK/3112/07.03.00/2021) and the Finnish Registry for Kidney Diseases (permission based on the meeting minutes dated 4 July 2019). The biobank access decisions for FinnGen samples and data used in FinnGen Data Freeze 10 include approvals from the following biobanks: THL Biobank (BB2017_55, BB2017_111, BB2018_19, BB_2018_34, BB_2018_67, BB2018_71, BB2019_7, BB2019_8, BB2019_26, BB2020_1 and BB2021_65); Finnish Red Cross Blood Service Biobank (7 December 2017); Helsinki Biobank (HUS/359/2017, HUS/248/2020, HUS/150/2022 §§12–18 and §23); Auria Biobank (AB17-5154 and amendment 1 (17 August 2020), amendments BB_2021-0140, BB_2021-0156 (26 August 2021, 2 February 2022), BB_2021-0169, BB_2021-0179, BB_2021-0161, AB20-5926 and amendment 1 (23 April 2020) with its modification (22 September 2021)); Biobank Borealis of Northern Finland (2017_1013, 2021_5010, 2021_5018, 2021_5015, 2021_5023, 2021_5017 and 2022_6001); Biobank of Eastern Finland (1186/2018 and amendments §§22/2020, 53/2021, 13/2022, 14/2022 and 15/2022); Finnish Clinical Biobank Tampere (MH0004 and amendments (21 February 2020 and 6 October 2020), §§8/2021, 9/2022, 10/2022, 12/2022, 20/2022, 21/2022, 22/2022 and 23/2022); Central Finland Biobank (1-2017); Terveystalo Biobank (STB 2018001 and amendment dated 25 August 2020); Finnish Hematological Registry and Clinical Biobank (decision dated 18 June 2021) and Arctic Biobank (P0844: ARC_2021_1001).
Ethics approval for the UK Biobank study was obtained from the North West Centre for Research Ethics Committee (11/NW/0382). The UK Biobank data used in this study were obtained under approved application 78537.
The activities of the EstBB are regulated by the Human Genes Research Act, which was adopted in 2000 specifically for the operations of the EstBB. Individual-level data analysis in the EstBB was carried out under ethical approval 1.1-12/624 from the Estonian Committee on Bioethics and Human Research (Estonian Ministry of Social Affairs), using data according to release application S22, document 6-7/GI/16259 from the EstBB.
Study setup
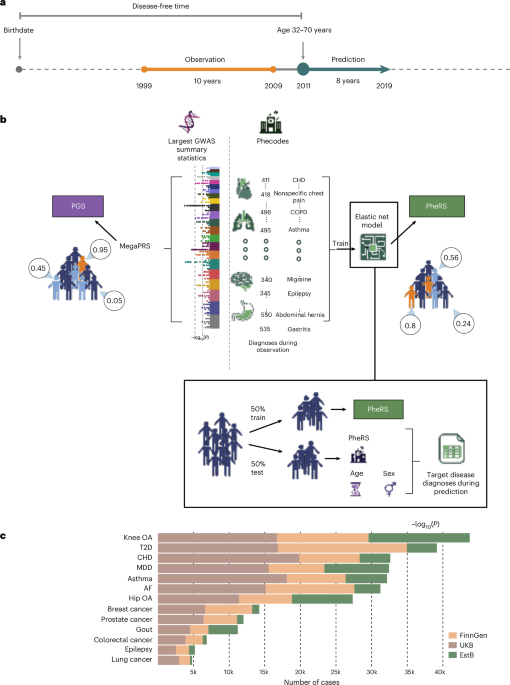
As outlined in Fig. 1b, each study consisted of a 10-year observation (6 years for EstBB due to shorter follow-up) and an 8-year prediction period, separated by a 2-year washout period. Each disease’s case and control definitions were based on diagnoses acquired in the 8-year prediction period (from 1 January 2011 to after 1 January 2019). The International Classification of Diseases (ICD) codes used to define the cases for each disease were based on previous harmonization between FinnGen and the EstBB phenotypes by the INTERVENE consortium34 (Supplementary Table 14). We consider all individuals as controls who were not cases. We only considered adults aged 32–70 in 1 January 2011 and removed all individuals diagnosed with the disease before this time. The lower limit for age of inclusion was chosen due to the inclusion of education level in some of the models and was determined based on the median age of obtaining a doctoral degree in the FinnGen dataset. Using this lower limit, most individuals included have finished their highest level of education. Furthermore, we removed all individuals with a diagnosis outside the prediction period (from 1 January 2011 to after 1 January 2019) and those lost to follow-up before the start of the prediction period. The ICD-codes used to define the cases for each disease and the number of cases and controls in each study are listed in Supplementary Tables 2 and 14.
We included 845,929 individuals (Supplementary Table 1) from three biobank-based studies—FinnGen29, UKB28 and EstB30 linked with national registers or EHRs. In FinnGen, we used Data Freeze 10, which includes 412,090 individuals, of whom 266,179 were aged 32–70 years in 1 January 2011. The longitudinal ICD-code diagnoses used to define the phecodes and the case and control status for each disease were based on in- and outpatient hospital register information. The UKB study included 464,076 individuals aged 40–70 years, with the ICD-code diagnoses based on inpatient information. The EstB study included 199,868 individuals, of whom 115,674 were aged 32–70 years. Here we also had primary care data as well as self-reported diagnoses available. More details on the phenotype harmonization can be found in ref. 34 and the Supplementary Methods.
PredictorsPGS
The PGS were previously computed by the INTERVENE consortium34 and based on the recent publicly available GWAS summary statistics, with minimal overlap with our study cohorts (Supplementary Table 15) using MegaPRS51 with the BLD-LDAK heritability model. For the Cox-PH models, we removed individuals from the studies that were part of the GWAS on which the PGS were based. Due to the large overlap with the UKB individuals, we only had PGS for gout, epilepsy, breast and prostate cancer available in the UKB.
PheRS
For the EHR-based models, we trained elastic net models32 on ICD-9 and ICD-10 diagnoses mapped to phecodes. The phecode mapping was based on v1.2b1 of the phecode map33,35 from https://phewascatalog.org/, with some manual additions. Since we only considered diagnoses during the observation period starting in 1999, all diagnoses were ICD-10-based in our data. To obtain the most comprehensive mapping, we removed all special characters from the ICD codes. If a match could not be found in the phecode map, we shortened the code by one digit until it could be mapped or was removed. The complete mapping used can be found in Supplementary Table 16. We gathered all phecodes in their three-digit parent node in the phecode ontology; for example, type 1 diabetes (250.1), T2D (250.2), and T2D with ketoacidosis (250.21) were all mapped to the same phecode diabetes mellitus (250). For each disease, we separately excluded predictors that were part of the exclusion range of the phecodes (Supplementary Table 3), for example, for T2D, we did not use secondary diabetes (phecode 249), diabetes mellitus (250) and conditions complicating pregnancy (649) as predictors. The phecode conditions complicating pregnancy was excluded because it was the parent node of the phecode Diabetes or abnormal glucose tolerance complicating pregnancy (649.1), which is in the exclusion range of the phecode for T2D (250.2). We only considered phecodes with a prevalence of at least 1% of the study population (Supplementary Table 11).
We implemented the PheRS using the LogisticRegression function from scikit-learn (version 1.3.2)52. We included age (at the start of the prediction period 1 January 2011) and sex as predictors in the PheRS models because they are important predictors, and otherwise the models would reconstruct predictors for age and sex using combinations of the phecode diagnoses, which would make interpretation of the phecode coefficient values challenging. Nonetheless, the effect of age and sex was then regressed out when evaluating the performances of the PheRS (see below). Models were penalized with the elastic net penalty. Predictors were coded as 1/0, where 1 = ‘predictor observed during the observation window’ and 0 = ‘predictor not observed during the observation window’, for each disease separately. For training, 50% of the data was used, and this was further divided into training (85%) and hold-out test (15%) sets. Sizes of the training datasets are shown for each disease and study in Supplementary Table 22. L1 to L2 ratio hyperparameter of the elastic net models was optimized using grid search and fivefold cross-validation over the range 0.05–0.95 (step size = 0.05), simultaneously with inverse of the regularization strength (C) over the following possible values: 1 × 10−5, 5 × 10−5, 1 × 10−4, 5 × 10−4, 1 × 10−3, 5 × 10−3, 1 × 10−2, 5 × 10−2, 1 × 10−1, 5 × 10−1, 1. Balanced class weights were used, based on class frequencies in the training data. The LOO analysis assessing the impact of the removal of individual phecodes to PheRS performance was performed using a ridge penalty instead of elastic net. This was done to cut running time substantially, as using ridge removes the L1 to L2 ratio hyperparameter and its optimization. Otherwise, the ridge models were fitted similarly to the elastic net models. Before running the LOO analysis, we tested that switching to ridge did not generally reduce the PheRS performance in FinnGen (Extended Data Fig. 7a).
Model fitting was done using stochastic average gradient descent. The best L1 to L2 ratio was selected based on the average precision score using 5-fold cross-validation on the training split. Missing values of predictors were imputed to the mean of the corresponding predictor in the study-specific training data, and all predictors were standardized to zero mean and unit variance on the study-specific training data before model fitting.
The PheRS models trained within the UKB or the EstB data on 50% of individuals were used to make predictions in FinnGen and UKB test sets, as is without any retraining within the studies. Standardization and imputation were performed based on the biobank-specific training data, meaning that, for example, when assessing the performance of the UKB-trained model in FinnGen, the FinnGen test set data were imputed and standardized based on the feature-specific means and s.d. from the UKB.
Cox-PH models
Ultimately, each individual was assigned 13 different PGS and PheRS scores describing their risk of getting a disease diagnosis in the prediction period based on genetic or EHR-based information. To make the PheRS and PGS comparable, we regressed out the effect of age, sex and the first ten genetic PCs from all continuous scores using the residuals from a logistic regression with the score as outcome. When only considering PheRS performance, we regressed out only age and sex. Subsequently, we scaled all predictors to have a mean of zero and a s.d. of 1. We then used these scores in separate Cox-PH analyses, with survival time defined as the period from 2011 until diagnosis, censoring (end of follow-up), or the end of the prediction period.
Additionally, we considered the CCI (Supplementary Methods)37,38—developed to account for the individual’s overall comorbidity burden—and the individual’s highest achieved education level in 2011 as an indicator of their socioeconomic status. For the CCI, we compared the top 10% of individuals with the highest CCI to the rest. The high-risk group included individuals with a CCI ≥ 2 and a few younger ones with a CCI of 1. For the highest education level, we mapped each study’s education coding to the 2011 International Standard Classification of Education (ISCED-11; Supplementary Table 17) codes. We compared the risk of individuals with basic education (ISCED-11: 1–4) to those who achieved higher education levels (ISCED-11: 5–7).
Statistics
We used the survival53 package (version 3.2-7) in R for creating the Cox-PH models and the Hmisc54 package (version 5.1.0) to calculate the c indices and 95% CIs. For a Cox-PH model with binary outcomes, the predicted survival times can be shown to be equal to the survival probability, so the c index is equivalent to the area under the receiver operating characteristic curve (AUC)55,56. The meta-analysis of the HRs and c indices was performed using the metafor57,58 package (version 4.6-0) in R with a random effects model. We used two-tailed P values, calculated using the pnorm function in the stats package (version 3.6.2) in R, based on the z scores of the β differences to compare the differences in HR magnitudes and one-tailed P values for the statistical testing of increases in the c indices. Additionally, we used Bonferroni correction to account for multiple hypothesis testing in each study (n = 13). Correlations were calculated, using the cor.test function from the stats package in R. For regressing out the covariates—age, sex and PCs—from the PheRS and PGS, we used scaled residuals from glm models with the stats package in R.
Comparison of phecode coefficients between different PheRS models
The elastic net hyperparameters were separately optimized for each PheRS model. This means that the absolute magnitudes of the coefficients for phecodes are not comparable between different PheRS. However, the relative importances of phecodes can still be compared, that is, whether, for example, the same phecodes are among the most important predictors in two different PheRS. To make visualization of the phecode importances in different PheRS clearer, we standardized the coefficients of each PheRS separately to a mean of 0 and a s.d. of 1 for the display items. Further, in each study, we ranked the phecodes in descending order by the PheRS coefficient values and assigned them ascending ranks. Thus, a lower rank indicates a higher PheRS coefficient in the model. Both the unscaled PheRS coefficients and ranks are Supplementary Table 12.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.