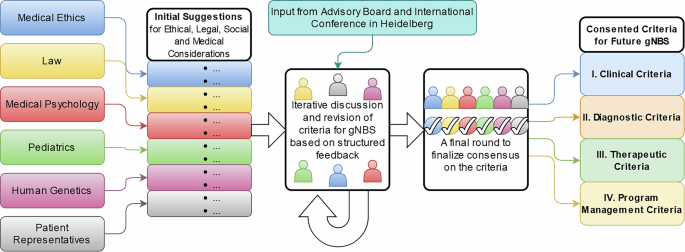
The above-described methodology resulted in a multi-dimensional framework for a gNBS program consisting of 18 screening criteria (Table 1) assigned to two overarching categories and four subcategories:
(A)
Criteria that enable transparent disease selection:
I.
Four clinical criteria,
II.
Four diagnostic criteria, and
III.
Three therapeutic-interventional criteria
(B)
Criteria to establish, manage, and further develop the gNBS program:
IV.
Seven program management criteria.
Table 1 Criteria for a genomic newborn screening program (by the project group NEW_LIVES).
A target disease qualifies for inclusion in a population-wide gNBS program if all screening criteria in subcategories I–III are met, while any (g)NBS program should meet all seven program management criteria (subcategory IV) if established on a population-wide level.
I. Clinical criteria (Characteristics of the target disease)
(1)
The gene-disease association of the target disease is “definitive” or “strong” according to the ClinGen classification.
Evidence for the causal relationship between a gene and a disease should have been repeatedly demonstrated in both research and clinical diagnostics (“definitive”) or reported independently in ≥ 2 studies (“strong”). The expert panel recommends the use of a Gene-Disease Validity Classification Information, such as curated by ClinGen [27]. For gene-disease associations not covered by ClinGen, the same standards as in ClinGen should apply.
(2)
The penetrance of variants associated with the target disease is at least 80% for the corresponding gene.
Only target diseases with high penetrance based on moderate to substantial evidence according to ClinGen [28] should be considered. For genetic diseases with multiple symptoms, penetrance refers to the clinically relevant main symptom. For target diseases not covered by ClinGen, the same standards as in ClinGen should apply. Target diseases with a penetrance of at least 80% would be, e.g., Nephropathic cystinosis (OMIM #606272) or Retinoblastoma (OMIM #614041).
(3)
The target disease is severe. This means that if left untreated, it results
(a)
in premature death, or
(b)
in major morbidity and a significantly impaired quality of life, or
(c)
in modest morbidity with a significantly impaired physical or cognitive development.
For the assessment of the severity, the ClinGen classification system or a comparable database can be used.
The expert panel has decided not to exclusively refer to the ClinGen classification as it leaves considerable room for interpretation. Diseases with “modest morbidity”, such as moderate intellectual disability, may not generally meet the definition of a severe disease, but are considered severe if associated with significant impairment of physical or cognitive development.
(4)
The average age of onset of the target disease is < 7 years of age.
To avoid waiting periods until disease-onset and as applicable for most current NBS programs’ target diseases, only diseases with an average age of onset in early childhood should be included. The specific age limit of under seven years was chosen because children can give assent or dissent to genetic testing themselves from this age onwards. As genetic diseases usually have a wide range of age of onset, the average should be considered to trade-off between the benefit for children with earlier onset and avoidance of “patients in waiting” [29]. This is particularly important as any inclusion of a disease into NBS will lead to expansion of the known clinical phenotype, especially towards attenuated phenotypes, which are likely to shift the age of onset to a later timepoint [1].
II. Diagnostic criteria (Requirements of the test)
(5)
A gNBS has proven advantages compared to the established diagnostic methods for the target disease. This means the target disease (1) cannot be detected by analytic methods of current NBS, but (2) can be detected by gNBS (a) more reliably (i.e., with higher sensitivity or specificity) or (b) at an earlier timepoint than with established methods of routine diagnostics.
A population-wide gNBS is only justified if identified patients have an additional benefit compared to identification by current NBS of asymptomatic newborns or established diagnostic methods used for symptomatic individuals. Consequently, gNBS is not intended to replace current NBS but to be added for diseases that cannot be identified by current screening methods (#5 (1)) if the described advantages (#5 (2)) exist.
(6)
The target disease can be clearly identified by molecular genetics using genome sequencing. The technical quality parameters of the test correspond to those of accredited diagnostic laboratories. The gene variants to be reported are detected with an analytical-diagnostic sensitivity and specificity of almost 100%.
The quality parameters of the test must meet the highest diagnostic standards and (likely) pathogenic variants must be detected. However, incomplete clinical sensitivity, caused by a high proportion of variants of uncertain significance (VUS), some of which may later be classified as pathogenic, should not be considered a general exclusion criterion for a target disease: if not all affected individuals receive a genetic diagnosis, those with a (likely) pathogenic variant in this gene would still benefit from reporting the findings. Genome sequencing can be used to identify single nucleotide variants, small rearrangements, and copy number variations, while, e.g., methylation defects cannot be identified with genome-sequencing methods commonly used today. This must be considered when selecting target diseases and adapted to new technologies.
(7)
Only “likely pathogenic” and “pathogenic” variants (ACMG classification) are reported. Carrier status and VUS are not reported.
The American College of Medical Genetics and Genomics’ (ACMG) criteria for interpretation of sequence variants recommend five categories for classifying the probability that the variant is disease causing: pathogenic, likely pathogenic, uncertain significance (VUS), likely benign, and benign [30]. Information on symptoms and family history is required for classification, which is not available in the screening context. Therefore, and to achieve the highest possible specificity, only variants of the first two categories should be reported. Autosomal recessive heterozygous variants not relevant for the newborn’s health should not be reported. In X-linked inheritance, hemizygous males (karyotype 46, XY) are affected by the disease. Heterozygous females (46, XX) are usually less severely affected or unaffected; exceptions exist. Therefore, (likely) pathogenic variants in target diseases with X-linked inheritance are usually only reported in males.
(8)
The suspected diagnosis from gNBS is confirmed by (1) a molecular genetic test from an independently obtained sample (e.g. new blood sample) or (2) an established non-genetic diagnostic test (e.g. biochemical, enzymatic, image morphological, histological, electrophysiological).
Screening tests differ from diagnostic tests. They identify individuals at risk for the identified condition, preferably during the preclinical stage of the disease. Thus, positive screening tests always require a careful and reliable confirmatory strategy, particularly, as in rare diseases false positive results are relatively frequent. For current NBS, biochemical and targeted genetic tests are used for confirmation [31]. Target diseases included in gNBS programs equally require a predefined pathway of suitable confirmatory tests. Whenever possible, confirmation diagnostics should include non-genetic functional biomarkers. If these are unavailable, confirmation should be based on a secondary genetic analysis with an alternative molecular genetic test in a new sample to exclude sample mix-up. A careful evaluation of the newborn’s phenotype and family history should be used to complete the confirmatory pathway. If the confirmatory diagnostics are based on a second genetic test, the biallelic phasing of the variants should be confirmed in autosomal recessive inheritance. If there are medically equivalent ways of confirmatory diagnostics, the one with the least discomfort for the child should be chosen.
III. Therapeutic-interventional criteria (Prerequisites of the intervention)
(9)
A therapeutic intervention is established and available that demonstrably has a beneficial effect on the course of the target disease, which means that it prevents, alleviates, or delays the onset of disease-specific signs and symptoms. If this requires recurrent monitoring, interventions in the form of regular surveillance examinations are also established and available.
Other possible benefits, such as non-medical actionability, secondary benefits to parents, siblings, or society, family planning implications, avoidance of a “diagnostic odyssey”, or just the wish to know, are not considered sufficient to justify the inclusion of a target disease in a population-wide NBS or gNBS.
An ongoing clinical trial would not suffice to consider an intervention as “established”.
“Available” means that all screened individuals have equal access to treatment and specialized teams offering monitoring and treatment. If gNBS also included diseases that did not require therapeutic intervention from the time of diagnosis but recurrent surveillance, e.g., RB1-related retinoblastoma, serial examinations necessary for regular surveillance as well as therapeutic interventions in the event of disease development would have to be established and available.
(10)
Pre- or early symptomatic start of intervention after identification of the target disease through gNBS is feasible and has a proven health benefit compared to starting the intervention after diagnosis by established methods of routine diagnostics.
If pre- or early symptomatic initiation of therapy did not provide a health benefit, post-symptomatic targeted routine diagnostics would be sufficient and the candidate disease would not qualify for inclusion as target disease in gNBS. Since genome sequencing is frequently performed at the appearance of first symptoms and even ultra-rapid genome sequencing is increasingly available for critically ill children [32], the sole prevention of a “diagnostic odyssey” does not justify the inclusion into gNBS.
(11)
The benefit of the intervention clearly outweighs its risks and burdens for the child. The risk is low or moderate, comparable to a classification of the intervention risk of ≥ 2 (“low risk”; “moderately acceptable risk”) according to ClinGen. The assessment of benefits, risks and burdens considers the effects of the intervention on the child’s quality of life.
The known risk and burden of therapeutic interventions or surveillance examinations should be outweighed by their potential benefit. Since the assessment of risks and benefits includes normative components, patient representatives should be involved in the process alongside medical and medical-ethics experts. Priority should always be given to the quality of life of the affected child because the burden and risk of an intervention as classified in ClinGen [33] may not (always) correspond to the perception of those affected [34].
IV. Program management criteria (Structure of the program)
Any NBS, including a gNBS, should be regarded as a public health measure. To offer guidance for the process of establishing, evaluating, and maintaining a (g)NBS program, seven criteria for program management have been defined (##12–18; Table 1).
Equal access (#12), an important ethical requirement of any public health program, comprises guarantee of cost coverage for screening, confirmatory diagnostics, and consecutive interventions and care. Current NBS programs generally provide equal access to all newborns in the target region, and enable a very high participation rate, e.g., about 99.9% in Germany [35]. In contrast, cost coverage for recommended therapies is still incomplete, e.g., special diets are not regularly paid for by health insurances [36]. Nevertheless, achieving the greatest possible equality, including full-cost coverage of recommended follow-up care, should be the aim of any (g)NBS program.
An additional challenge for equality are minorities for whom only limited population-specific genetic data might be available for genomic analysis. Since precise knowledge of the genetic variants in the screened population is an essential condition for the success of a gNBS, it seems sensible to include underrepresented populations in registry studies in order to achieve equitable representation.
While equality of access (#12) implies that every newborn’s guardians are offered gNBS, participation needs to remain voluntary (##13–14). Although written informed consent (IC) is not uniformly required across all NBS programs worldwide [37], it should be mandatory for gNBS as for any genetic diagnostic test. Since gNBS is a complex topic, there is a balance to be achieved between information overload and insufficient information (#14). Withdrawal from gNBS is possible at any time (#14f) with the exception that a positive gNBS result, once obtained, will be communicated in the interest of the child.
To make gNBS as minimally invasive for the child as possible, the same dried blood spot sample should be used for NBS and gNBS, utilizing the established system of filter paper cards. Sample collection, transport, and analysis (#15), confirmation and communication of test results, and the initiation of recommended interventions (#16) should follow clearly defined guidelines including exact procedures for each target disease. It is essential for a gNBS program that precise information about the procedure following a positive result, including specific contact persons, is provided to families directly after a positive screening report.
The use of data, and samples from gNBS requires strict adherence to data protection laws (#17). Samples, and data from gNBS, as well as clinical follow-up data are of great value for program evaluation, and quality assurance (#17a–d), but also for adjustment, and improvement of the program. In addition, they might contribute to research on genetic diseases, potentially resulting in new or improved treatment, and to individual benefits. Since genetic data is considered particularly sensitive, it requires special protection. Therefore, the expert panel recommends that such secondary use of data should only be made possible either with separate written IC or where legally permitted (#17e), in both cases with additional safeguards (#17f), in compliance with applicable law and regularly updated to reflect changing legal requirements.
A gNBS should be organized as an integrated and learning public health program with central coordination, data-driven evaluation of quality, safety and acceptability, and re-evaluation of target diseases, including case definitions and recommended interventions (#18).